BtGal27E (exo-α-Galactosidase)
BtGal27E
Ex-Gal0036
(EC.3.2.1.22)exo-α-Galactosidase
CAZy Family:GH27
PROPERTIES
1.ELECTROPHORETIC PURITY
-Single band on SDS-gel electrophoresis (MW ~69kDa)
Figure 1. Electrophoresis analysis ofBtGal27E. M, molecular weight marker (PageRuler Prestained Protein Ladder, Thermo Scientific); lane 1, culture lysate before IPTG induction; lane 2, culture lysate after IPTG induction; lane 3,BtGal27Epurified from Ni sepharose fastflow column.
2.SPECIFIC ACTIVITY
0.13 U/mg protein (on pNP-α-gal) at pH 7.0 and 30°C.
One Unit of pNP-α-gal activity is defined as the amount of enzyme required to release 1 μmol of p-nitrophenyl per minute from pNP-α-gal (5 mM) in phosphate buffer (20 mM) pH7.0.
3.RELATIVE RATES OF HYDROLYSIS OF SUBSTRATES
Table 1. Relative activity ofBtGal27Eon different substratesa.
Substrateb |
Relative activity (%)c |
pNPβGlu |
_ |
pNPβGal |
_ |
pNPβMan |
_ |
pNPβXyl |
_ |
pNPαGlc |
_ |
pNPαGal |
100±0.0 |
pNPαMan |
_ |
pNPαAraf |
_ |
pNPαArap |
_ |
aReactions were performed with 5 mM substrate, pH7.0, at 30°C for 10 min.
bAbsorption caused by released p-nitrophenol was measured at 405 nm. The relative activity on pNPαgal was taken as 100%.
cThe data are reported as means±standard errors from the mean for three independent experiments.
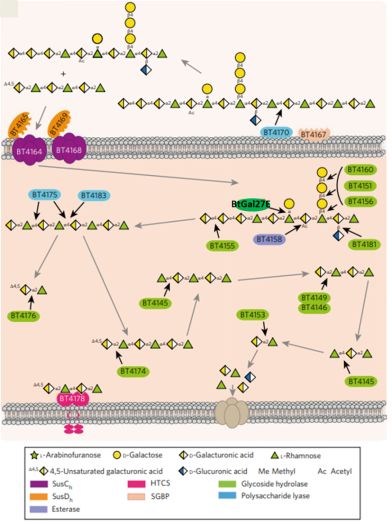
Figure 2. Model of pectin utilization by B. thetaiotaomicron. Model for degradation of rhamnogalacturonan I is displayed. The black arrows indicate the linkage cleaved by the various enzymes, while the grey arrows show the direction of the degradative pathway.
4.PHYSICOCHEMICAL PROPERTIES
pH Optima:7.0
Temperature Optima:37°C
5.STORAGE CONDITIONS
The enzyme should be stored at -20 °C. For assay, this enzyme should be diluted in phosphate buffer (20 mM) pH 7.0. Swirl to mix the enzyme immediately prior to use.
6. REFERENCES
[1]Cartmell A, Muñoz-Muñoz J, Briggs J A, et al. A surface endogalactanase in Bacteroides thetaiotaomicron confers keystone status for arabinogalactan degradation[J]. Nature microbiology, 2018, 3(11): 1314-1326


