CcBgl3B(exo-β-1,6-Glucosidase,CcGlu3A,CB-3B)
CcGlu3A(CcBgl3B, CB-3B)
Ex-Glu0010
(EC.3.2.1.10) exo-β-1,6-Glucosidase
CAZy Family: GH3
PROPERTIES
1.ELECTROPHORETIC PURITY
-Single band on SDS-gel electrophoresis (MW ~84 kDa)
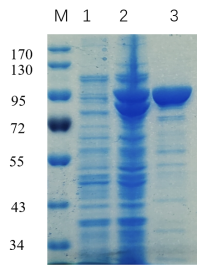
Figure 1. Electrophoresis analysis of CcGlu3A. M, molecular weight marker (PageRuler Prestained Protein Ladder, Thermo Scientific); lane 1, culture lysate before IPTG induction; lane 2, culture lysate after IPTG induction; lane 3, CcGlu3A purified from Ni sepharose fastflow column.
2.SPECIFIC ACTIVITY
26.5 U/mg protein (on pNP-β-glu) at pH 6.0 and 30oC
One Unit of pNP-β-glu activity is defined as the amount of enzyme required to release 1 μmol of glucose per minute from pNP-β-glu (5 mM) in phosphate buffer (50 mM) pH 6.0.
3.RELATIVE RATES OF HYDROLYSIS OF SUBSTRATES
Table 1. Relative activity of CcGlu1C on different substratesa.
Substrate |
Relative activity (±SDb) |
pNPβGlc |
100±0.0 |
pNPβGal |
5.1±0.6 |
pNPβMan |
_ |
pNPβXyl |
_ |
pNPαGlc |
_ |
pNPαGal |
_ |
pNPαMan |
_ |
pNPαAraf |
_ |
pNPαArap |
_ |
Sophorose |
25.1±0.7 |
Laminaribose |
40.0±1.3 |
Cellobiose |
_ |
Gentiobiose |
100±0.0 |
aReactions were performed with 1 mM (p-nitrophenyl glycosides and disaccharides) or 5 mg/ml (polysaccharides) substrate, pH 6.0, at 30oC for 10 min.
bThe data are reported as means ± standard errors from the mean for three independent experiments.
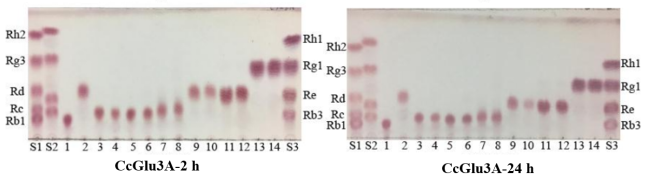
Figure2. Analysis of ginsenosidase activity of CcBgl1C by TLC. S1, standards of Rh2, Rg3, Rd, Rc and Rb1; S2, standards of CK, Gyp LXXV, Gyp XVII and Rb2; 1, biotransformation substrate Rb1; 2, transformed products of Rb1;3, biotransformation substrate Rb2; 4, transformed products of Rb2; 5, biotransformation substrate Rb3; 6, transformed products of Rb3; 7, biotransformation substrate Rc; 8, transformed products of Rc; 9, biotransformation substrate Rd ; 10, transformed products of Rd; 11, biotransformation substrate Re ; 12, transformed products of Re; 13, biotransformation substrate Rg1; 14, transformed products of Rg1; S3, standards of Rh1, Rg1, Re and Rb3.
4.PHYSICOCHEMICAL PROPERTIES
pH Optima: 6.0
pH Stability: 6.0-9.0
Temperature Optima: 55°C
Temperature Stability:<45°C
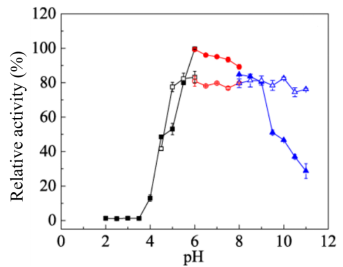
Fig. 3. Effect of pH on activity (solid symbols) and stability (hollow symbols) of CcGlu3A. The optimal pH of CcBgl3A was determined by ranging pH from 2.0 to 11.0 using following buffers: sodium acetate buffer, pH 2.0–6.0; Na2HPO4–NaH2PO4buffer, pH 6.0–8.0; Glycine–NaOH buffer, pH 8.0–11.0. The maximum activity obtained was defined as 100%. The pH stability of CcGlu3A was determined by pre-incubating CcGlu3A in different pH for 24 h at 4°C and then determining the percentage of residual activity under standard assay conditions. The activity of CcGlu3A without pre-incubating was defined as 100%. Results are presented as means ± standard deviations (n = 3).
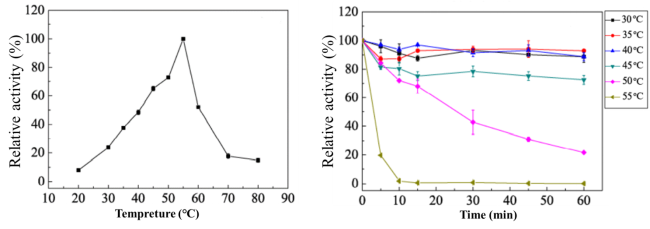
Fig. 4. Effect of temperature on activity (a) and stability (b) ofCcGlu3Ausing pNP-Glu as substrate. The optimal temperature (a) was determined at different temperatures from 20 to 80oC. The maximum activity obtained was defined as 100% activity. Thermal stability was determined by incubating the enzyme for 1 h at different temperatures. The activity of the enzyme before incubation was defined as 100%. Results are presented as means ± standard deviations(n = 3).
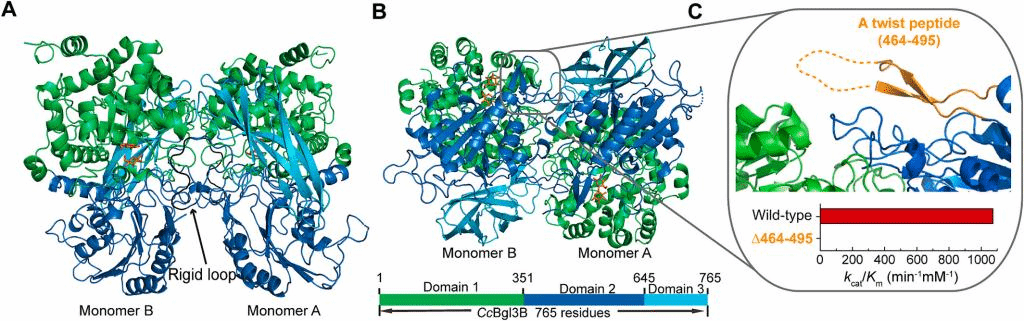
Fig. 5. Overall costructure of CcBgl3B with carbohydrates. Domains 1 (residues 3–351), 2 (residues 352–645) and 3 (residues 646–765) of CcBgl3B are shown in green, marine and cyan, respectively. (A) Top view of the CcBgl3B dimer. The CcBgl3B dimer (PDB: 8XRX) is composed of monomers A and B. The gentiobiose bound to monomer A and the enzyme-glycosyl intermediate in monomer B are shown (carbon, orange; oxygen, red). The rigid loop (residues 622–630) is shown in black. (B) Side view of the CcBgl3B dimer. (C) The “forehead” of CcBgl3B. (For interpretation of the references to colour in this figure legend, the reader is referred to the web version of this article.)
5.STORAGE CONDITIONS
The enzyme should be stored at -20°C. For assay, this enzyme should be diluted in phosphate buffer (50 mM) pH 6.0. Swirl to mix the enzyme immediately prior to use.
6. REFERENCES
[1] Hu C, Yuan Y, et al. A trapped covalent intermediate as a key catalytic element in the hydrolysis of a GH3 β-glucosidase: An X-ray crystallographic and biochemical study. International Journal of Biological Macromolecules. 2024, 265, 131131


